1 Overview
The Delta Science Program is once again partnering with the National Center for Ecological Analysis and Synthesis (NCEAS) to convene a collaborative data synthesis working group in the spring of 2026. Delta Synthesis Working Groups receive advanced data science and statistics training with immediate opportunities to use those newly acquired skills to analyze available data and produce relevant research and data products. The upcoming working group will be the third cohort held through this partnership.
2 Learning Objectives
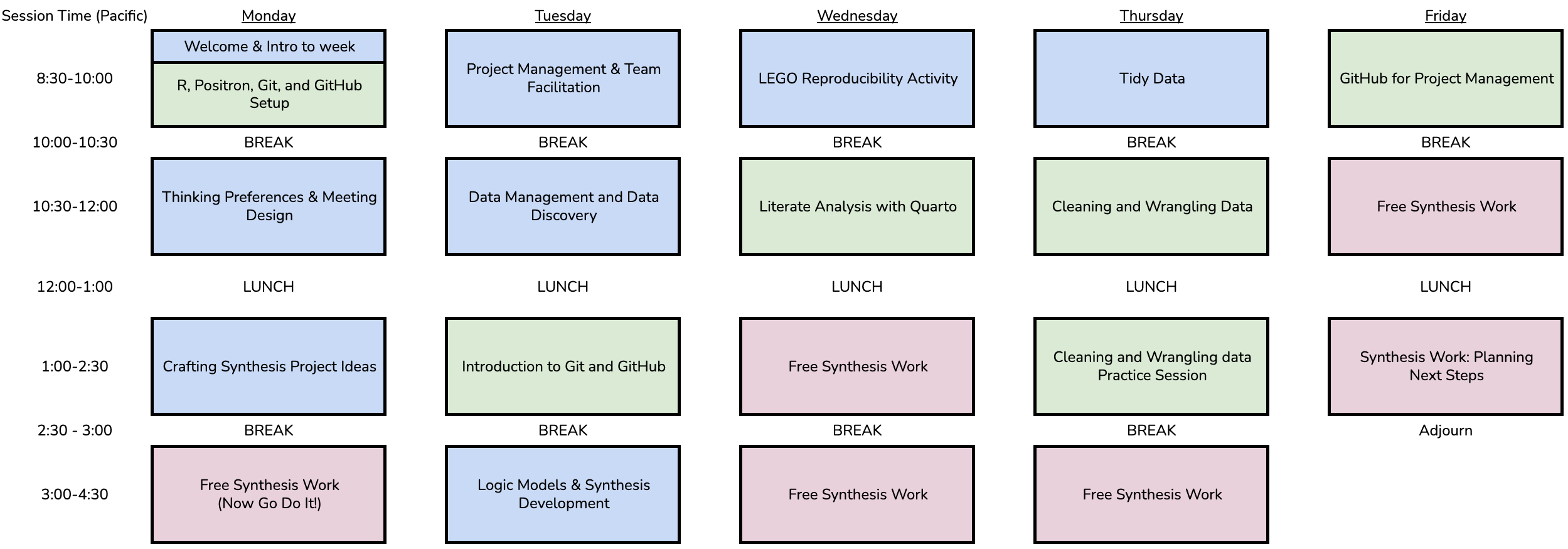
3 Course Schedule
Table Color Legend: Technical Topic; Non-Technical Topic; Synthesis Work
| Day | Time | Module Name | Primary Instructor |
|---|---|---|---|
| 1 | 8:30 - 9a | Welcome & Intro to Week | - |
| 1 | 9 - 10a | R, Positron, Git, and GitHub Setup |
Jim Regetz; Casey O’Hara; Nick Lyon |
| 1 | 10 - 10:30a | Break | - |
| 1 | 10:30 - 12p | Thinking Preferences & Meeting Design |
Nick Lyon |
| 1 | 12 - 1p | Lunch | - |
| 1 | 1 - 2:30p | Crafting Synthesis Project Ideas | Nick Lyon |
| 1 | 2:30 - 3p | Break | - |
| 1 | 3 - 4:30p | Free Synthesis Work (Now Go Do It!) |
- |
| Day | Time | Module Name | Primary Instructor |
|---|---|---|---|
| 2 | 8:30 - 10a | Project Management & Team Facilitation |
Nick Lyon |
| 2 | 10 - 10:30a | Break | - |
| 2 | 10:30 - 12p | Data Management & Data Discovery |
Jim Regetz |
| 2 | 12 - 1p | Lunch | - |
| 2 | 1 - 2:30p | Introduction to Git and GitHub | Casey O’Hara |
| 2 | 2:30 - 3p | Break | - |
| 2 | 3 - 4:30p | Logic Models & Synthesis Development |
Casey O’Hara |
* Tentatively choose your group by EoD
| Day | Time | Module Name | Primary Instructor |
|---|---|---|---|
| 3 | 8:30 - 10a | LEGO Reproducibility Activity | Jim Regetz; Casey O’Hara; Nick Lyon |
| 3 | 10 - 10:30a | Break | - |
| 3 | 10:30 - 12p | Literate Analysis with Quarto | Casey O’Hara |
| 3 | 12 - 1p | Lunch | - |
| 3 | 1 - 2:30p | Free Synthesis Work | - |
| 3 | 2:30 - 3p | Break | - |
| 3 | 3 - 4:30p | Free Synthesis Work | - |
| Day | Time | Module Name | Primary Instructor |
|---|---|---|---|
| 4 | 8:30 - 10a | Tidy Data | Jim Regetz |
| 4 | 10 - 10:30a | Break | - |
| 4 | 10:30 - 12p | Cleaning and Wrangling Data | Jim Regetz |
| 4 | 12 - 1p | Lunch | - |
| 4 | 1 - 2:30p | Cleaning and Wrangling Data Practice Session |
Jim Regetz; Casey O’Hara; Nick Lyon |
| 4 | 2:30 - 3p | Break | - |
| 4 | 3 - 4:30p | Free Synthesis Work | - |
| Day | Time | Module Name | Primary Instructor |
|---|---|---|---|
| 5 | 8:30 - 10a | GitHub for Project Management | Jim Regetz |
| 5 | 10 - 10:30a | Break | - |
| 5 | 10:30 - 12p | Free Synthesis Work | - |
| 5 | 12 - 1p | Lunch | - |
| 5 | 1 - 2:30p | Synthesis Work: Planning Next Steps | - |
| 5 | 2:30 - 3p | Adjourn | - |
4 Pre-Workshop Preparation
If you are participating in this workshop series, we are so excited to help you acquire new or hone existing technical and interpersonal skills in the realm of synthesis science! To ensure that you get the most out of this workshop series, please follow the instructions on the Pre-Workshop Prep page before the first week.
5 Code of Conduct
By participating in this activity you agree to abide by the NCEAS Code of Conduct.
6 About this book
These written materials are the result of a continuous and collaborative effort at NCEAS to help researchers make their work more transparent and reproducible. This work began in the early 2000’s, and reflects the expertise and diligence of many, many individuals. The primary authors are listed in the citation below, with additional contributors recognized for their role in developing previous iterations of these or similar materials.
This work is licensed under a Creative Commons Attribution 4.0 International License.
Citation: Casey O’Hara, Nick J. Lyon & Jim Regetz (2026), NCEAS Synthesis Skills Training for the Delta Science Program, April 2026, NCEAS Learning Hub. nceas-learning-hub.github.io/2026_delta_week1.
Content contributors: Ben Bolker, Amber E. Budden, Julien Brun, Angel Chen, Samantha Csik, Halina Do-Linh, Marty Downs, Sarah Elmendorf, Natasha Haycock-Chavez, S. Jeanette Clark, Carrie Kappel, Li Kui, Julie Lowndes, Stephanie Hampton, Matt Jones, Samantha Katz, Nick J. Lyon, Gregory Maurer, Erin McLean, Bryce Mecum, Casey O’Hara, Deanna Pennington, Karthik Ram, Jim Regetz, Tracy Teal, Camila Vargas Poulsen, Daphne Virlar-Knight, Leah Wasser.
This is a Quarto-built website. To learn more about Quarto sites visit https://quarto.org/docs/websites/.